Nitrogen metabolism and regulatory networks in corynebacteria
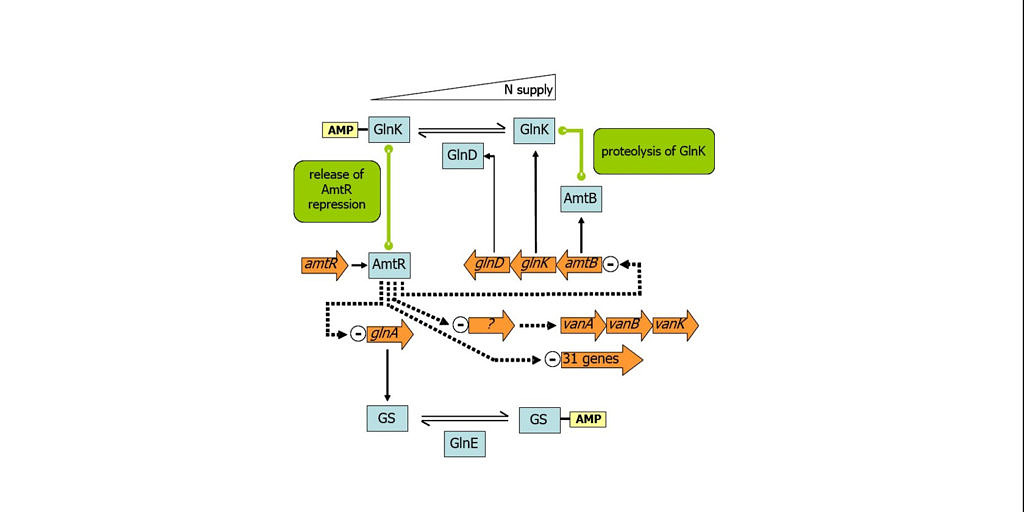
Nitrogen is an essential component of nearly all of the complex macromolecules in a bacterial cell, e.g. proteins, nucleic acids and cell wall components. Consequently, most prokaryotes have developed elaborate control mechanisms to provide an optimal supply of nitrogen for growth and to cope with situations of nitrogen limitation. We are interested in the uptake of nitrogen sources, their assimilation and connected regulatory systems in Corynebacterium glutamicum, a Gram-positive soil bacterium used for the industrial production of amino acids. The core adaptations of C. glutamicum to nitrogen limitation on the level of transcription are controlled by the global regulator AmtR. Further components of the signal pathway are GlnK, a PII-type signal transduction protein, and GlnD, encoding an adenylyltransferase regulating GlnK function. Mechanisms involved in nitrogen control in C. glutamicum regulating gene expression and protein activity are repression of transcription, protein-complex formation, protein modification by adenylylation, change of intracellular localization and proteolysis.